Project Overview - Beta
A single-cell eQTL atlas of the developing human brain
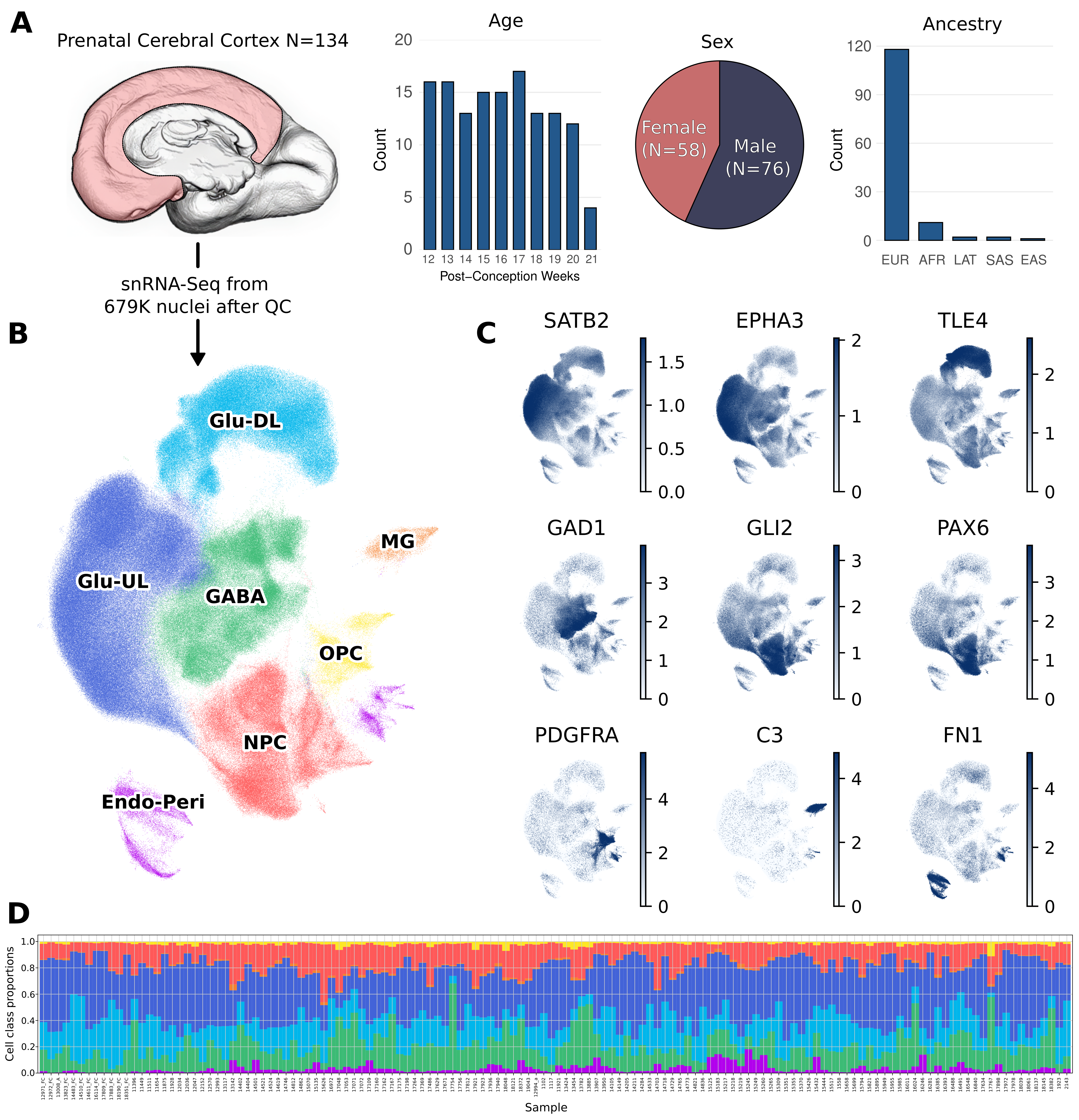
We performed single-nucleus RNA sequencing and genome-wide genotyping on cerebral cortex from 134 unrelated samples (second trimester) to generate the first cell-type-resolved eQTL atlas of the prenatal human brain.
This site is the documetation for an end-to-end computational genomics platform to process ~3 TB of raw single-nucleus RNA sequencing and genome-wide genotyping data. The pipeline identifies genetic variants that influence gene expression in specific brain cell types during development, and links those variants to neuropsychiatric disease risk.
NOTE: the documentation and eQTL browser app are currently in beta.
Data Engineering Architecture
This project processes ~3 TB of raw genomic data through a suite of 13 interoperable Snakemake pipelines, orchestrated on a SLURM HPC cluster. The platform was designed from the ground up for scalability, reproducibility, and collaborative reuse.
Workflow Orchestration
All pipelines are managed by Snakemake (v8.x) with a dedicated SLURM cluster profile, allowing up to 500 concurrent jobs across a dedicated compute partition (c_compute_neuro1, account scw1641). The profile handles automatic job submission, logging, and resource allocation per rule:
# config/profile/config.yaml (excerpt)
executor: cluster-generic
jobs: 500
use-conda: true
use-singularity: true
cluster-generic-submit-cmd: >
sbatch
--ntasks={resources.ntasks}
--mem={resources.mem_mb}
--time={resources.time}
--cpus-per-task={resources.threads}
--account=scw1641
--partition=c_compute_neuro1The pipeline is launched via a single shell script that captures the full Snakemake log and emails on completion:
# workflow/snakemake.sh
snakemake --profile ../config/profile/ $@ 2> smk-"`date +"%d-%m-%Y"`.log
mail -s "Snakemake has finished" camerond@cardiff.ac.uk < smk-"`date +"%d-%m-%Y"`.logCentralised Configuration
All parameters, file paths, tool settings, container paths, GWAS URLs, and cell-type lists are managed in a single config/config.yaml. This means the entire platform can be reconfigured for a new dataset by editing one file — no hardcoded paths in any script or rule.
Key config-driven components include:
- Cell types (20 entries: 7 broad + 12 subtypes) — propagated automatically to TensorQTL, SuSiE, S-LDSR, SMR, and TWAS rules via Snakemake wildcards
- GWAS URLs — six neuropsychiatric GWAS (SCZ, BPD, MDD, ADHD, OCD) downloaded directly from Figshare/PGC by the pipeline
- Container paths — eight Singularity containers mapped centrally, ensuring every rule uses the correct software environment
- Analysis parameters — SuSiE window (1 Mb), batch count (25), FDR threshold (0.05), TensorQTL permutation bounds, SMR windows, all in one place
Data Ingestion: JSON-driven FASTQ Management
Raw sequencing data from three plates (150 samples, 43 sublibraries) are spread across multiple sequencing runs and lanes on network-attached storage. Rather than hardcoding paths, we use a custom Python script (workflow/scripts/create_parse_json.py) to crawl FASTQ directories, match files by sample and read orientation using regex, sort across lanes and runs, and serialise the result to a JSON manifest:
# Matches filename pattern: 10_S8_L001_R1_001.fastq.gz
m = re.search(r'(\d+)_S\d+_(L\d{3})_(R\d)', file)
if m:
sample, lane, reads = m.group(1), m.group(2), m.group(3)
FILES[sample][reads].append(full_path)The resulting JSON (e.g. config/samples_plate3.json) maps each sample ID to its full list of R1 and R2 FASTQ paths across all lanes and runs. Snakemake ingests this at runtime via a lambda wildcard function, allowing it to concatenate the correct files for each sample dynamically:
rule cat_fqs:
input:
r1 = lambda wildcards: MERGE_FQ[wildcards.sample]['R1'],
r2 = lambda wildcards: MERGE_FQ[wildcards.sample]['R2']A parallel JSON (config/bam_files.json) maps sample IDs to their processed BAM file paths for downstream genotype-aware steps (cellSNP-lite, Vireo donor deconvolution).
Reproducible Environments
Software reproducibility is enforced at two layers:
Conda environments with fully pinned dependencies are used for Python-based pipelines (Scanpy, TensorQTL). The eqtl_study.yml environment pins every package to an exact build hash, ensuring bit-for-bit reproducibility across HPC nodes and future reruns.
Singularity containers are used for R-based and specialist tools where conda environments are insufficient. Eight containers are defined in config.yaml:
| Container | Purpose |
|---|---|
tensorqtl.sif |
GPU-accelerated eQTL mapping (PyTorch) |
r_eqtl.sif |
Core R analysis environment |
susier_v24.01.1.sif |
SuSiE fine-mapping |
seurat5f.sif |
Seurat 5 / general R |
twas.sif |
FUSION TWAS weight computation |
gtex_eqtl.sif |
FastQTL / GTEx tools |
genotype-qc2hrc_latest.sif |
Genotype QC and TOPMED imputation prep |
ubuntu_22.04.sif |
Lightweight shell utility container |
GPU-accelerated eQTL Mapping
TensorQTL (PyTorch backend) is used for cis eQTL mapping, enabling GPU-parallelised permutation testing across all 19 cell types. Four mapping modes are run per cell type: nominal, permutation, independent, and trans. Output is stored in Parquet format for efficient downstream parsing.
Automated Documentation
Pipeline documentation (this site) is built with Quarto and published automatically to GitHub Pages via a GitHub Actions workflow on every push to main:
# .github/workflows/publish.yml
- name: Render and Publish
uses: quarto-dev/quarto-actions/publish@v2
with:
target: gh-pagesPipeline Overview
The 13 pipelines run in a defined order, passing outputs directly between stages:
| # | Pipeline | Input | Output |
|---|---|---|---|
| 01 | Parse Alignment | Raw FASTQs | Per-plate DGE matrices (H5AD) |
| 02 | Scanpy | H5AD matrices | Annotated clusters, pseudobulk counts |
| 03 | Genotypes (pre-imputation) | Raw PLINK files (hg19) | TOPMED-ready VCFs (hg38) |
| 04 | Genotypes (post-imputation) | Imputed VCFs | Filtered, annotated, indexed VCF |
| 05 | TensorQTL | Pseudobulk counts + VCF | cis eQTL results (nominal, perm, indep, trans) |
| 06 | eQTL Replication | eQTL results | π₁ enrichment vs. 4 public datasets |
| 07 | Prep GWAS | PGC summary statistics | Munged, lifted, harmonised GWAS files |
| 08 | SuSiE Fine-mapping | eQTL + VCF | Credible sets, MaxCPP / CS95 annotations |
| 09 | S-LDSR | Fine-mapped eQTLs + GWAS | Partitioned heritability results |
| 10 | SMR | eQTL summary stats + GWAS | Colocalisation results |
| 11 | TWAS Weights | Pseudobulk counts + genotypes | FUSION weight files per gene/cell type |
| 12 | cTWAS | TWAS weights + GWAS | Causal TWAS results |
| 13 | Visualisation | All upstream results | Manuscript figures and tables |
Repository
Full source code, configuration, and documentation are available at: github.com/Dazcam/eQTL_study_2025